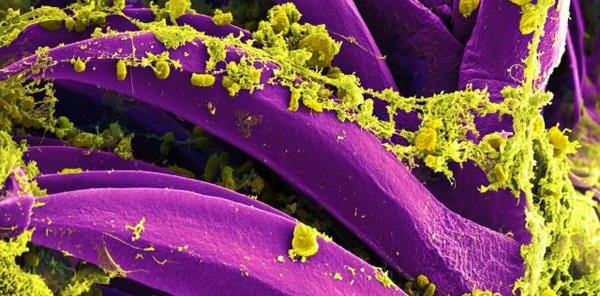
The mere mention of the plague brings to mind the devastating “Black Death” pandemic that spread across Europe in the 1300s. Mass graves were piled high with the corpses of its millions of victims, while the disease rampaged across Europe for many decades. Yersinia pestis, the bacterium responsible for that plague pandemic, still persists in the environment among rodent and flea populations today, and human outbreaks regularly occur around the world. Most recently, an outbreak of plague was confirmed late last year in Madagascar as well as within a prairie dog colony in Colorado just this June.

CDC, CC BY
Y. pestis can cause three different forms of plague: bubonic, pneumonic and septicemic. Pneumonic plague infects the lungs, causing severe pneumonia. It’s the most serious form of the disease, with fatality rates approaching 100% if untreated, although recovery is possible with antibiotics if caught in time. While increased basic hygiene and developments in modern medicine have greatly reduced the severity of plague outbreaks, the symptoms of pneumonic plague are so similar to that of the flu that misdiagnosis or delays in treatment can have fatal consequences.
Y. pestis is known to have evolved from the relatively mild gut pathogen Yersinia pseudotuberculosis sometime within the last 5,000 to 10,000 years – very recently on an evolutionary timescale. Sometime during this evolution Y. pestis developed new modes of transmission and disease manifestations, which allowed it to adapt to new animals and environments. Rather than simply causing an upset stomach, the bacterium became the killer we know from the Middle Ages.

Kamal Kishore / Reuters
One of our lab’s major research goals is to figure out how Y. pestis developed its ability to specifically cause pneumonic plague. Our research, recently published in Nature Communications, offers new insights into how small genetic changes fundamentally affected the emergence of Y. pestis as a severe respiratory pathogen.
Prior to our study, the consensus in the field has been that pneumonic plague was a secondary byproduct of the invasive disease associated with bubonic plague. As pneumonic plague represents only 5%–10% of current plague infections in humans, the field has presumed that pneumonic plague occurs only once Y. pestis reaches the lungs following systemic infection, as might occur during bubonic plague. While this may be the case now, it may not necessarily represent what occurred in the past, especially as Y. pestis was just emerging from its ancestor Y. pseudotuberculosis.

CDC/ Dr Jack Poland, CC BY
First, target the lungs
Therefore, we began our study by asking a relatively simple question: “When did Y. pestis develop the ability to infect the lung and cause pneumonic plague?” Remember, it was only recently, evolutionarily speaking, that it started targeting the lungs rather than the gut. Y. pestis is believed to have emerged as a species 5,000–10,000 years ago, but the first known pandemic of plague in humans didn’t occur until the Justinian Plague that afflicted the Byzantine empire about 1,500 years ago.

Museum of London, Schuenemann et al PNAS vol. 108 no. 38
A recent discovery helped us investigate. Scientists successfully recovered DNA from Y. pestis from human skeletons in a Black Death mass grave in London, England. The genetic material from the historic site is very similar to DNA isolated from recent modern plague outbreaks. The fact that the DNA from then is similar to the DNA from now indicates that today’s Y. pestis has maintained its devastating disease-causing capability.
To answer the question of how Y. pestis made that crucial leap to targeting the lung and therefore being able to cause pneumonic plague, we used strains of both ancestral and modern Y. pestis in our study. These ancestral strains of Y. pestis, isolated from voles in the Transcauscaian highland, carry characteristics of both modern, pandemic Y. pestis and the relatively benign predecessor species Y. pseudotuberculosis that still exists today.
Thus, these ancestral versions can be considered “intermediate” strains, trapped somewhere between the gut Yersiniae and modern, virulent Y. pestis. Indeed, these “intermediate” lineage ancestral strains are as closely related to Y. pseudotuberculosis as we can get while still technically representing species of Y. pestis. Because of their unique genetic characteristics, these ancestral strains can provide crucial insights into how this bacterium may have adapted to new host environments as it evolved from Y. pseudotuberculosis.
Surprisingly, we found that these ancestral strains were able to cause pneumonic plague in a manner indistinguishable from that of modern Y. pestis in mice – but only if the bacteria carried the gene for a single protein called Pla. Pla is unique to Y. pestis and was acquired very early in the evolution of the species.
Almost all ancestral strains of Y. pestis carry the gene for Pla, but there still exist a few that represent ancestral Y. pestis just prior to acquisition of Pla. We were able to test if these pre-Pla strains were able to cause pneumonic plague – and they did not. But as soon as Y. pestis picked up this gene, the bacteria could cause epidemics of pneumonic plague. No further changes were necessary, even though there are dozens of additional differences between these ancestral strains and modern Y. pestis. So Y. pestis was able to cause pneumonic plague much earlier in its history than had previously been thought – as soon as it acquired this single gene for Pla.

Justin Eddy, Lindsay Gielda, et al, CC BY-ND
Second, increase infectiousness
But that’s not where the story ends. It turns out that all modern pandemic strains of Y. pestis contain a single amino acid mutation in Pla compared to ancestral Y. pestis. This change slightly alters the function of the Pla protein. The mutation, however, plays no role in the ability of any Y. pestis isolates to cause pneumonic plague – ancestral or modern.
Quite surprisingly, this modification allowed the Y. pestis to spread deeper into host tissue following a bite from an infected flea or rodent, leading to the development of bubonic plague with its trademark swollen lymph nodes. This suggests that Y. pestis was first a respiratory pathogen before it was able to efficiently cause invasive infections.
This discovery challenges our traditional notion of how plague evolved. Rather than pneumonic plague being a late addition to Y. pestis’s arsenal as commonly believed, its ability to target the lung came before the change that makes it such an infectious pathogen. Our research suggests that the acquisition of Pla and its ability to cause pneumonic plague occurred well before 1,500–5,000 years ago. But the amino acid modification didn’t occur until just prior to 1,500 years ago, allowing Y. pestis to become much more deadly. All strains of Y. pestis from the time of the Justinian Plague and after have the deadly modification of Pla, while strains prior do not.

Paul Fürst
Our results may explain how, through one small amino acid change, Y. pestis quickly transitioned from causing only localized outbreaks of disease to the pandemic spread of Y. pestis as seen during the Justinian Plague and the Black Death.
And it raises the ominous possibility that other respiratory pathogens could emerge from similar small genetic changes.
![]()
Daniel Zimbler is Postdoctoral Fellow in Bacteriology at Northwestern University.
Wyndham Lathem is Assistant Professor of Microbiology-Immunology at Northwestern University.
This article was originally published on The Conversation.
Read the original article.