By Peter White
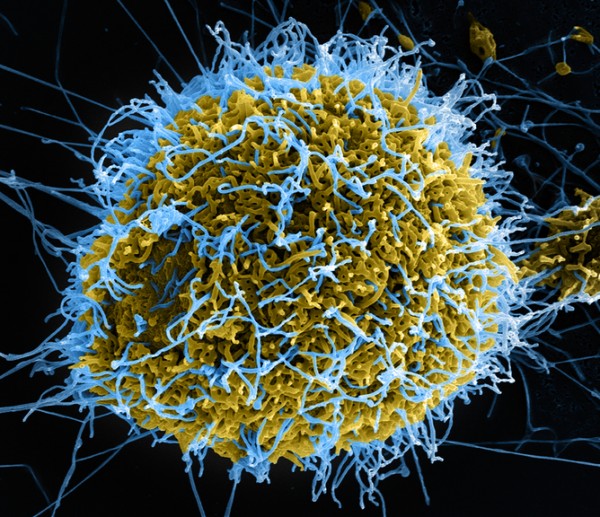
As the worst known epidemic of the Ebola virus continues in West Africa, scientists around the world are trying to develop treatments for those infected. But a process of viral mutation, known as “genetic drift”, could potentially compromise their efforts.
Genetic drift is one reason why RNA viruses, such as influenza and norovirus, cause global epidemics of disease (pandemics) about every three years. To understand how it works, we need to go back to basics.
Ebola is an RNA virus, the fastest-evolving genetic entity we know. These viruses continually change their genetic sequence because the enzymes responsible for making more copies of their genetic code often make mistakes. The process is akin to photocopying a photocopy: the image changes slowly over time. And these changes are known as genetic drift.
Genetic drift lets the new viruses change the surface protection proteins that the immune system targets. This enables them to infect people who have immunity against previous versions.
The main candidates
The World Health Organisation’s November 2014 report Potential Ebola Therapies and Vaccines lists four classes of medicines for Ebola: i) immunomodulators, ii) antiviral drugs, iii) immunoglobulins and iv) antiviral small inhibitory RNA (or RNA therapies).
Drugs in the latter two classes are clinically the most advanced, but are only in the earliest of the three stages of human clinical trials. These types of drugs are among the forerunners of a new wave of exciting RNA-based therapies. But they have not previously been widely commercialised for the treatment of viral infections.
Immunomodulators
The first class of drugs – immunomodulators – are designed to stimulate the innate immune response (which has broad antiviral effects) by triggering the production of many hundreds of antiviral proteins in infected cells.
When tested in rhesus macaques soon after an exposure to a lethal dose of the Ebola virus, this class of drug increased survival time, but failed to stop the monkeys from dying. Other treatments have resulted in better outcomes in animal models.
Immunomodulators are, so far, perhaps the weakest of the four classes of drugs that could be used against Ebola.
Antivirals
The most common type of antiviral drug is called a nucleoside analogue. Chemically, a nucleoside analogue “looks like” a sub-unit of RNA (a nucleotide), the building blocks of viral genetic codes. The virus mistakes this “rogue” building block while replicating – to its own demise. Further genome copying is shut down, reducing viral disease or even, in some cases, curing viral infections by eliminating the virus from the host.
Originally developed to fight influenza, T-705 or Favipiravir (Toyama Chemical) is a nucleoside analogue with broad antiviral activity. While its potency against Ebola is low compared to how well it works against influenza, results from the latest clinical trial show it’s ineffective for people who are very ill but works for those with low levels of the Ebola virus in their blood.
Another nucleoside analogue, known as BCX4430 (Biocryst), has protected macaque monkeys from Ebola-related Marburg virus infection and death, when administered two days after exposure.
RNA therapies
RNA drugs could be the next big thing to treat viral disease. The viral genetic code is translated into proteins through an intermediate called messenger RNA (mRNA). Once the cell is infected, the viral mRNA tells the cell to make viral proteins. By using small RNA molecules that match the viral mRNA sequence, there are a few ways to prevent the viral proteins from being made.

DFID/Flickr, CC BY
It’s analogous to taking out the middle man in a business arrangement. One partner makes some cakes, the middle man transports them to a shop and the third person sells the cakes. If you take out the driver, then no cakes get sold, or in the context of mRNA and Ebola, the viruses don’t replicate.
Tekmira Pharmaceuticals is developing a triple combination RNA-based drug for fighting Ebola, which is probably the most advanced. It has now been through Phase 1 human clinical trials. Another company, Sarepta Therapeutics, has used an alternative RNA therapy to target the same three Ebola messenger RNAs to prevent replication.
Both Sareota and Tekmira’s RNA-base drugs have successfully protected monkeys from dying after Ebola infection.
Immunoglobulins
Immunoglobulins, an antibody class of treatment, target a surface protein that viruses use to enter cells. It’s an obvious target for an antibody-based drug because when antibodies bind to this Ebola protein, the virus’ entry into cells is blocked and infection is prevented.
But viruses can mutate or mask their outer protein coats to evade host immune responses, so using a single antibody will eventually result in viral evasion. One way to counter this is to use a combination of antibodies to target many different parts of the viral protein coat. This reduces the viruses’ ability to escape and become resistant.
The drug ZMapp, which is an immunoglobulin, uses a cocktail of three antibodies to stop the viruses evading cells. It protects Ebola-infected monkeys from death and has been used on two people infected with the virus, who survived. Although it is not known whether they survived because of the drug, there are strong indications it has an impact.
Where to now?
Immunoglobulins and RNA therapies could both become less effective if Ebola mutates because they are based directly on viral genetic sequence. Both RNA therapies were designed against the original 1976 virus, while Zmapp was designed from the version of the Ebola virus circulating in 1995.
A recent paper evaluating the impact of genetic drift since these dates has found the Ebola genome has mutated by around 3% compared to both the 1976 and 1995 Ebola predecessors. This mutation rate is lower than for other RNA viruses, such as HIV, hepatitis C virus and influenza, but not by much. All RNA viruses mutate a lot and this poses a danger to the effectiveness of immunoglobulins and RNA therapies.
The good news is that genetic drift would be unlikely to result in viral resistance against nucleoside analogues. For this to occur, mutations within the “viral engine” (the polymerase gene), which copies the genome, are needed. And this type of mutations renders the virus weak compared to their non-mutated brethren. So a virus that could evade the drug through genetic drift wouldn’t be very virulent in the first place.
But drug resistance could occur in people who are treated with these medicines. That’s because the presence of the drug creates pressure for the virus to mutate its engine to survive as only resistant viruses will be able to replicate.
Genetic drift is unlikely to render newly developed drugs ineffective. The three most advanced therapies we have were formulated on older versions of the virus but their developers don’t seem concerned. This is because each company has three separate versions of the drug in its formula.
For the virus to become resistant, three viral mutations would need to occur simultaneously, and this is very difficult. Given what’s at stake here, we should be comforted by the fact that drug developers have taken genetic drift into account.
![]()
This article was originally published on The Conversation.
Read the original article.